CRISPR-Cas9 vectors for genome editing and host engineering in the baculovirus–insect cell system | PNAS

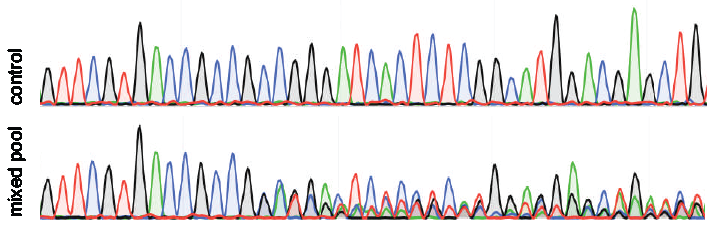
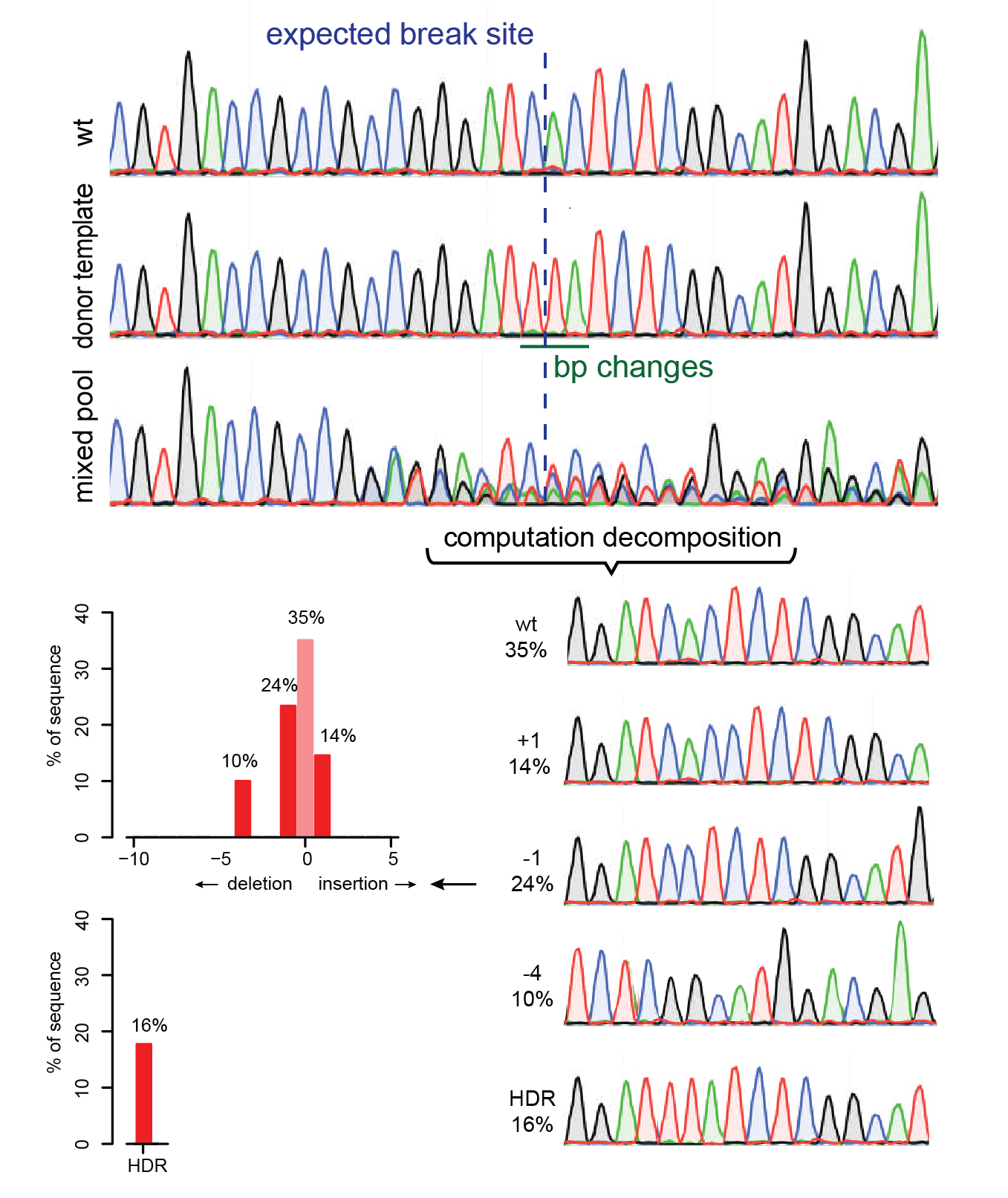
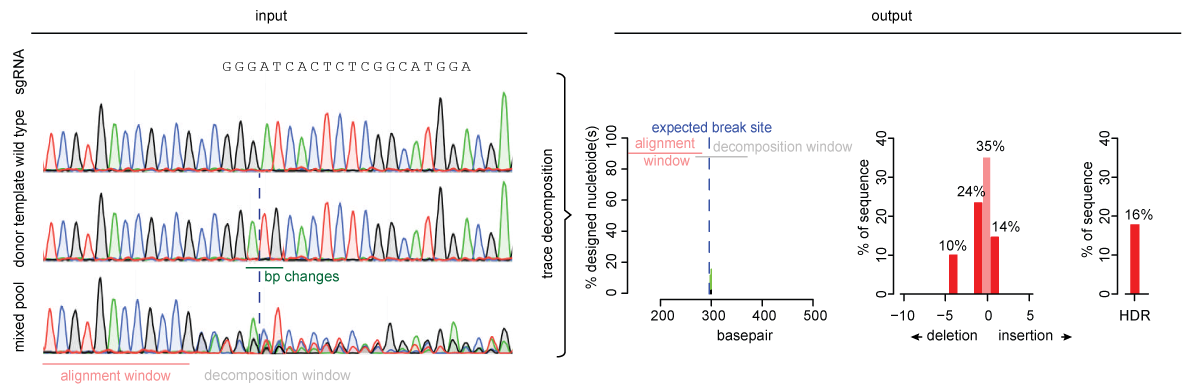
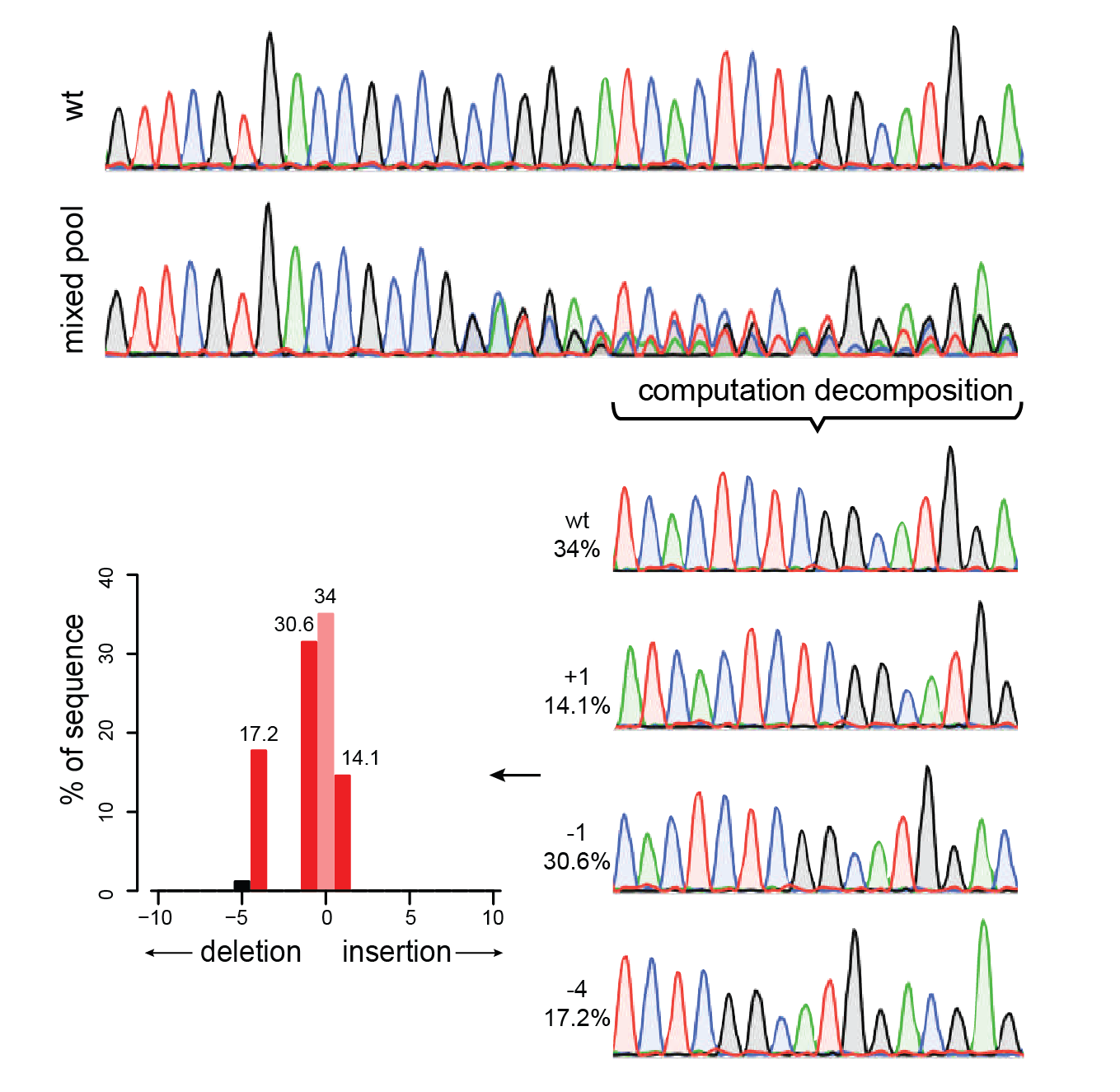
Analysis of gene modification conditions using TIDE software. The bar... | Download Scientific Diagram

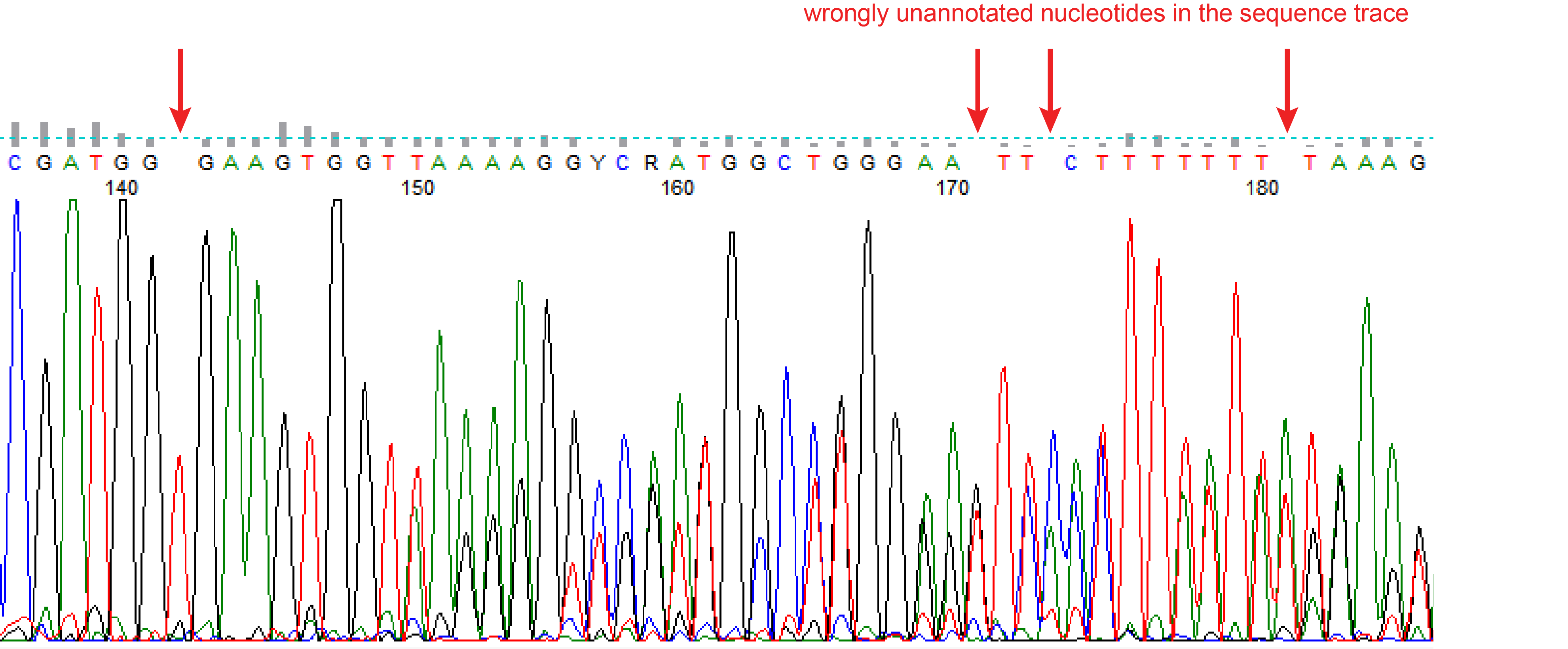
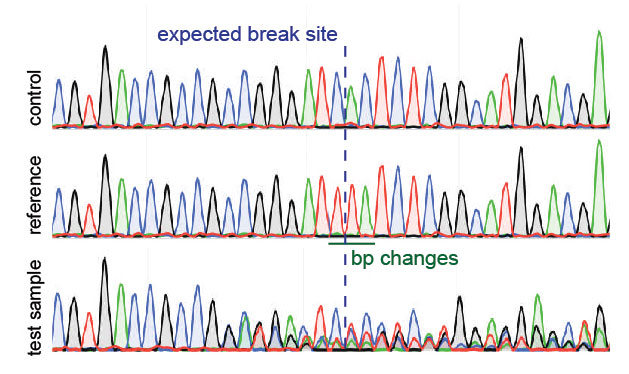
Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases - ScienceDirect

TIDE sequencing analysis of light-mediated indel mutation HEK293T cells... | Download Scientific Diagram
![Cells | Free Full-Text | A Versatile Strategy to Reduce UGA-Selenocysteine Recoding Efficiency of the Ribosome Using CRISPR-Cas9-Viral-Like-Particles Targeting Selenocysteine-tRNA[Ser]Sec Gene | HTML Cells | Free Full-Text | A Versatile Strategy to Reduce UGA-Selenocysteine Recoding Efficiency of the Ribosome Using CRISPR-Cas9-Viral-Like-Particles Targeting Selenocysteine-tRNA[Ser]Sec Gene | HTML](https://www.mdpi.com/cells/cells-08-00574/article_deploy/html/images/cells-08-00574-g004.png)
Cells | Free Full-Text | A Versatile Strategy to Reduce UGA-Selenocysteine Recoding Efficiency of the Ribosome Using CRISPR-Cas9-Viral-Like-Particles Targeting Selenocysteine-tRNA[Ser]Sec Gene | HTML
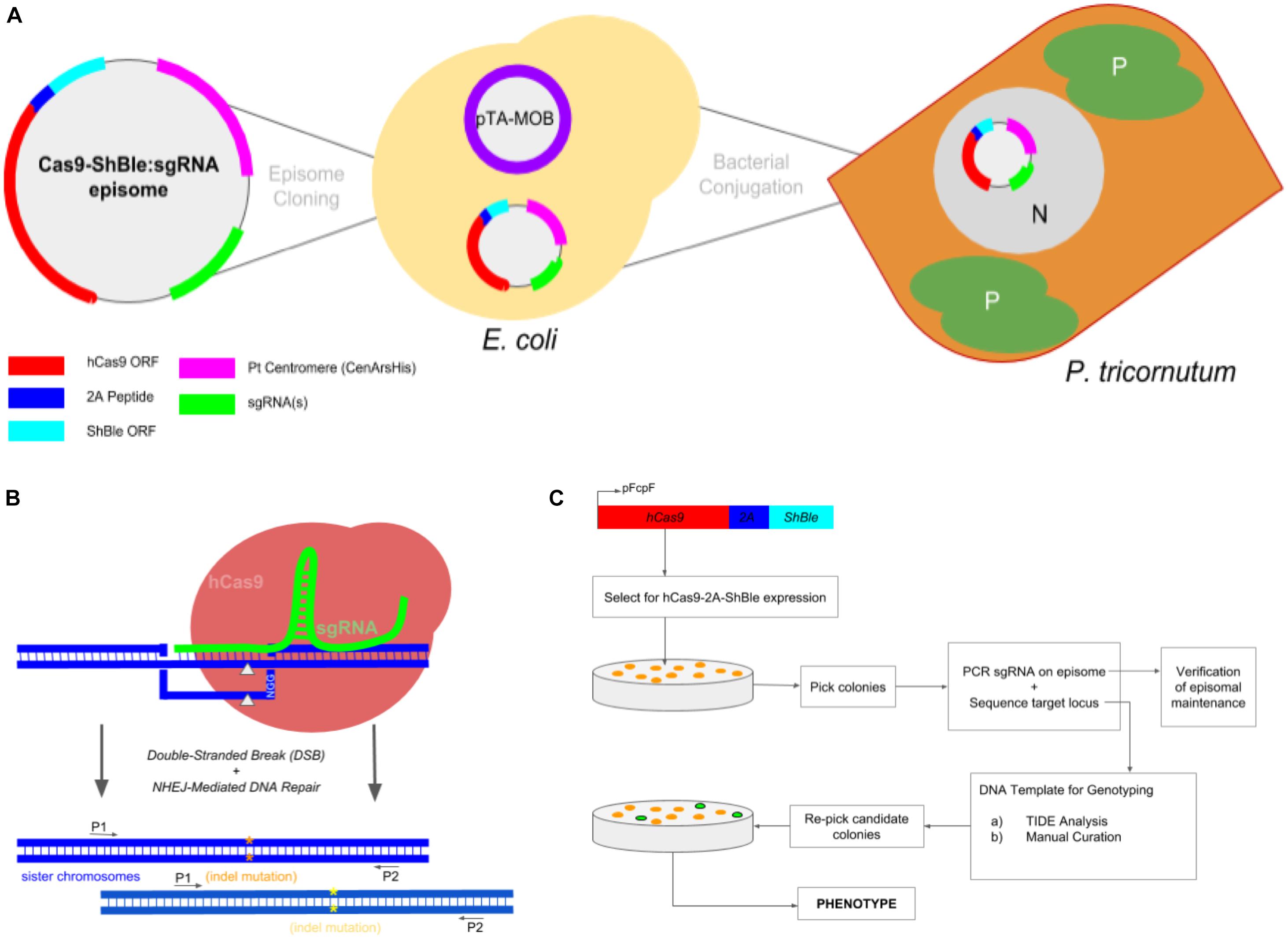
Frontiers | Multiplexed Knockouts in the Model Diatom Phaeodactylum by Episomal Delivery of a Selectable Cas9 | Microbiology

CRISPR/Cas9-Directed Reassignment of the GATA1 Initiation Codon in K562 Cells to Recapitulate AML in Down Syndrome: Molecular Therapy - Nucleic Acids
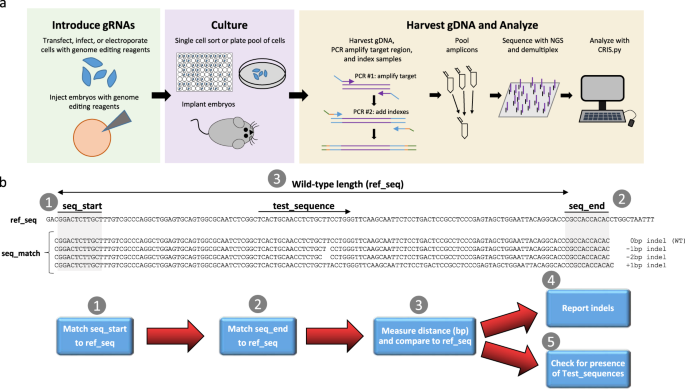
CRIS.py: A Versatile and High-throughput Analysis Program for CRISPR-based Genome Editing | Scientific Reports

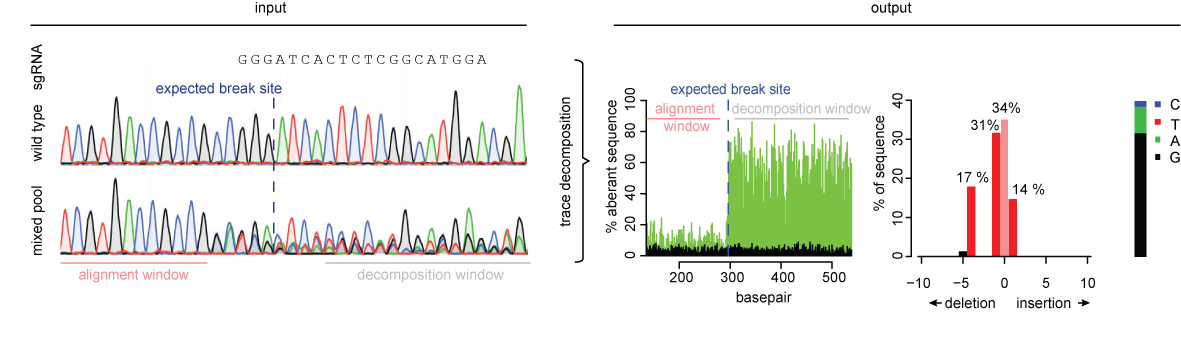
Netherlands Cancer Institute Grants Exclusive License to Desktop Genetics for TIDE | Technology Networks


![A dual sgRNA approach for functional genomics in Arabidopsis thaliana [OPEN] | bioRxiv A dual sgRNA approach for functional genomics in Arabidopsis thaliana [OPEN] | bioRxiv](https://www.biorxiv.org/content/biorxiv/early/2017/08/21/172676/F1.large.jpg)